 You can modify the columns that appear by clicking the column icon to the right of the gear wheel and checking the columns you want to appear in the table panel. The Table Panel on the lower right lets you see the attribute information for nodes, edges, and the network. Everything appears in one window, within different panels. Unlike Gephi, in Cytoscape there is no separate ‘Data Laboratory’ tab. Similarly to Gephi, the data will appear in the graph window as a grid: We have no column in blue because our edges for this dataset are not weighted (they do not have values assigned to them).ħ. Check the box next to ‘Show Text File Import Options’ under ‘Advanced’ to change the delimiter to ‘comma’ if your data is not formatted the way the table appears above.Ħ. In the section marked ‘Interaction Definition,’ select the appropriate columns so the color coding for Source Interaction, Interaction Type, and Target Interaction matches your data columns.ĥ. A window titled ‘Import Network From Table’ will appear. 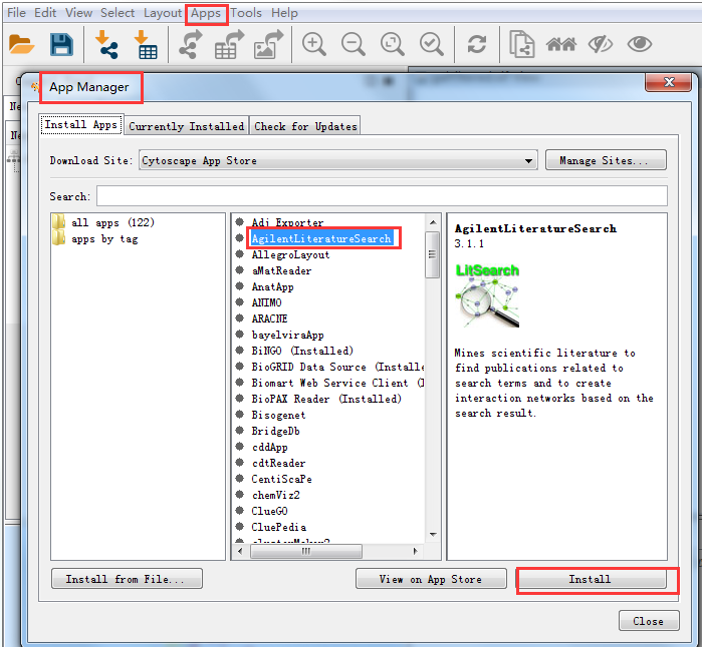
Go to File –>Import–>Network–>File and select the CSV file you created for the High Energy Arxiv Physicists’ dataset in the first part of the tutorial.Ĥ. Click OK for default selections in the ‘Create New Network’ window.ģ. Choose ‘Empty Network’ under ‘Start New Session’ in the Welcome Window.Ģ. 
To begin, go to and download version 3.2.1 of Cytoscape to your computer.Īfter running the installation, open the program:ġ. In this half of the tutorial, we’ll do the same for Cytoscape. The first post outlined preparing a dataset for upload into Gephi and covered how to get started with the styling options and layouts available in Gephi. This post is the second half of a two-part beginner’s introduction to network visualization. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
